The Institute of Bioinformatics and Medicine (IBM) of your country has been studying the DNA sequences of several organisms, including the human one. Before analyzing the DNA of an organism, the investigators must extract the DNA from the cells of the organism and decode it with a process called ``sequencing''.
A technique used to decode a DNA sequence is the ``shotgun sequencing''. This technique is a method applied to decode long DNA strands by cutting randomly many copies of the same strand to generate smaller fragments, which are sequenced reading the DNA bases (A, C, G and T) with a special machine, and re-assembled together using a special algorithm to build the entire sequence.
Normally, a DNA strand has many segments that repeat two or more times over the sequence (these segments are called ``repetitions''). The repetitions are not completely identified by the shotgun method because the re-assembling process is not able to differentiate two identical fragments that are substrings of two distinct repetitions.
The scientists of the institute decoded successfully the DNA sequences of numerous bacterias from the same family, with other method of sequencing (much more expensive than the shotgun process) that avoids the problem of repetitions. The biologists wonder if it was a waste of money the application of the other method because they believe there is not any large repeated fragment in the DNA of the bacterias of the family studied.
The biologists contacted you to write a program that, given a DNA strand, finds the largest substring that is repeated two or more times in the sequence.
The first line of the input contains an integer T
specifying the number of test cases (
1 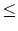 T
T 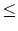 100
). Each test case consists of a single line of text that represents a DNA sequence S
of length n
(
1
100
). Each test case consists of a single line of text that represents a DNA sequence S
of length n
(
1 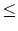 n
n 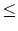 1000
).
1000
).
You can suppose that each sequence S
only contains the letters A, C, G and T.
For each sequence in the input, print a single line specifying the largest substring of S
that appears two or more times repeated in S
, followed by a space, and the number of ocurrences of the substring in S
.
If there are two or more substrings of maximal length that are repeated, you must choose the least according to the lexicographic order.
If there is no repetition in S
, print ``No repetitions found!''
6
GATTACA
GAGAGAG
GATTACAGATTACA
TGAC
TGTAC
TTGGAACC
A 3
GAGAG 2
GATTACA 2
No repetitions found!
T 2
A 2
Colombia'2008